 |
|
Contact Information
Office: MEYR 475D
Phone: 410-455-2512
Email: dkoirala@umbc.edu
|
Associate Professor
Professional Interests
RNA structures, functions and consequences
Research in our group focuses on understanding the structures and functions of RNA molecules involved in fundamental biological processes and exploring the opportunities for developing RNA-targeted therapeutics to treat both genetic and infectious diseases. Our laboratory interconnects biochemistry, biophysics and biology, providing the members of our laboratory an excellent opportunity to learn a wide array of both biochemical and biophysical techniques while pursuing cutting-edge research in the field of RNA structural biology. Followings are the current research directions in our laboratory.
RNA structures associated with cap-independent viral translation
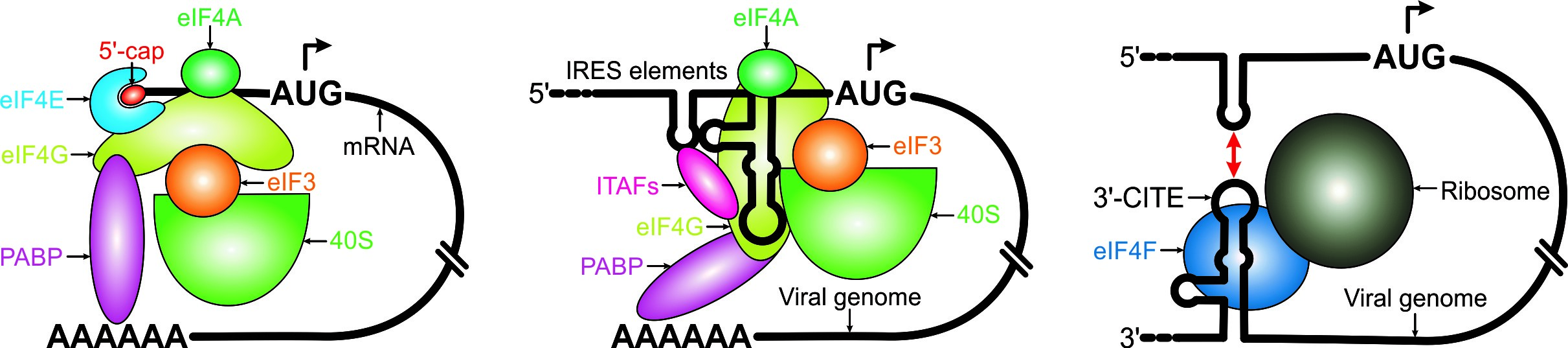
In contrast to the 5ꞌ-cap-based canonical translation in most eukaryotes, many viral genomes and a subset of cellular mRNAs are translated via cap-independent mechanisms that involve structured RNA elements such as internal ribosome entry sites (IRESs) and 3ꞌ cap-independent translation elements (3ꞌ-CITEs). However, our understanding on how these RNA structures are organized and how they recognize translation initiation factors or the ribosome remains largely elusive. Our laboratory is focused on determining the 3-dimensional structures of RNA elements associated with the viral cap-independent translation and understanding how these structures hijack the canonical, cap-dependent translation machinery from the host cell to mediate and regulate the non-canonical, cap-independent viral translation.
RNA structures associated with human repeat expansion disorders
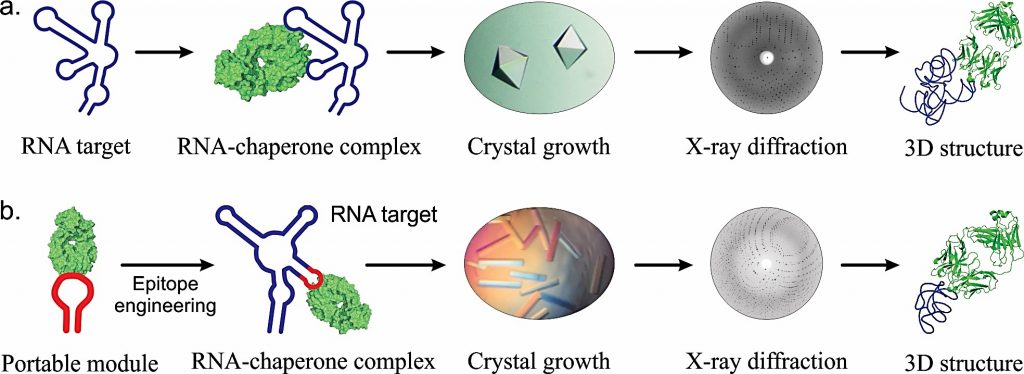
Besides the mechanisms that involve mutant proteins expressed from the mRNAs with expanded nucleotide repeats, pathogenic pathways in human disorders such as huntington’s disease, myotonic dystrophy fragile-X syndrome and amyotrophic lateral sclerosis also involve RNA-related toxicity induced by the RNA structures of the corresponding mRNAs. However, our understanding on the structures formed by these mRNAs and their specific roles in pathogenesis remains elusive. Our laboratory is focused on developing the strategies to study the structural elements in both protein-coding and non-coding regions of these mRNAs and their contributions to the disease pathogenesis. We use synthetic antibodies that specifically bind the RNA structures formed by the expanded mRNAs to investigate their structures, cellular locations and life-cycles.
Outcomes of our research will not only provide insights into the mechanisms of fundamental biological processes but also unlock ample opportunities for developing targeted therapeutics to treat both genetic and infectious diseases. Moreover, structural understanding on the landscape of RNA structures combined with mechanistic insights into how particular structural features enable the biological function will be tremendously useful for developing algorithms to predict new RNA structures using bioinformatics and computational tools.
Selected Publications
Das N.K., Vogt J, Patel A., Al Banna H., Koirala D., Structural basis for a highly conserved RNA-mediated enteroviral genome replication, Nucleic Acids Res., 2024, gkae627.
Ojha M., Vogt J., Das N.K., Redmond E., Singh K., Al Banna H., Sadat T., Koirala D., Structure of saguaro cactus virus 3′ translational enhancer mimics 5′ cap for eIF4E binding, Proceedings of the National Academy of Sciences, 2024, 121 (4), e2313677121.
Al Banna, H., Das, N. K., Ojha, M., Koirala D. Advances in chaperone-assisted RNA crystallography using synthetic antibodies. BBA advances, 2023, 100101.
Das, N. K., Hollmann, N. M., Vogt, J., Sevdalis, S. E., Banna, H. A., Ojha, M., Koirala D., Crystal structure of a highly conserved enteroviral 5′ cloverleaf RNA replication element. Nat. Commun., 2023, 14(1), 1955.
Krochmal D., Shao Y., Li N.S., DasGupta S., Shelke S.A., Koirala D., Piccirilli J.A., Structural basis for substrate binding and catalysis by a self-alkylating ribozyme, Nat. Chem. Biol., 2022, 18, 376–384 (co-corresponding author).
Koirala D., Lewicka A., Koldobskaya Y., Huang H. & Piccirilli J.A., Synthetic antibody binding to a preorganized RNA domain of hepatitis C virus internal ribosome entry site inhibits translation, ACS Chem. Biol., 15, 1, 205-216, 2020.
Koirala D., Shao Y., Koldobskaya Y., Fuller J. R., Watkins A.M., Shelke S.A., Pilipenko E.V., Das R., Rice P.A. & Piccirilli J. A., A conserved RNA structural motif for organizing topology within picornaviral internal ribosome entry sites, Nat. Commun., 10:3629, 2019
Koirala D., Shelke S.A., Dupont M., Ruiz S., Dasgupta S., Bailey L.J., Benner S.A. & Piccirilli J.A., Affinity maturation of a portable Fab-RNA module for chaperone-assisted RNA crystallography, Nucleic Acids Res., 46(5), 2624 – 2635, 2018
Koirala D., Shrestha P., Emura T., Hidaka K., Mandal S., Masayuki E., Sugiyama H. & Mao H., Single-molecule mechanochemical sensing using DNA origami nanostructures, Angew. Chem. Int. Ed. Engl., 2014, 53, 8137 – 8141
Koirala D., Punnoose J.A., Shrestha P. & Mao H., Yoctoliter thermometry for single-molecule investigations: a generic bead-on-a-tip temperature-control module, Angew. Chem. Int. Ed. Engl., 2014, 53, 3470 – 3474
Koirala D., Ghimire C., Bohrer C., Sannohe Y., Sugiyama H. & Mao H., Long-loop G-quadruplexes are misfolded population minorities with fast transition kinetics in human telomeric sequences, J. Am. Chem. Soc., 2013, 135, 2235 – 2241
Koirala D, Dhakal S., Ashbridge B., Sannohe Y., Rodriguez R., Sugiyama S., Balasubramanian S. & Mao H., A single-molecule platform for investigation of interactions between G-quadruplexes and small-molecule ligands, Nat. Chem., 2011, 3, 782-787
Honors and Awards
CAREER Award, National Science Foundation (NSF) (2023)
Collaborative Development Award, CRNA (2023)
Strategic Awards for Research Transitions (START), UMBC (2021)
Summer Research Faculty Fellowship (SURFF), UMBC (2021)
Taylor Scholarship for Graduate Research, Kent State University (2013)
GSS Domestic Travel Grant, Kent State University (2013)
University Fellowship, Kent State University (2012)
Pandit Narayan Prasad Upadhaya Award, Nepal Chemical Society(2006)
P.M. Singh Gold Medal, Tribhuvan University, Nepal (2005)
Amrit Gold Medal, Tribhuvan University, Nepal (2005)
Kirti-Sanu Gold Medal, Shesh-Hira Pratisthan, Nepal (2005)
Harihar Raj Lohani Award, Nepal Chemical Society (2003)
University Scholarship, Tribhuvan University, Nepal (1999-2004)
Courses Teaching
CHEM 437 Comprehensive Biochemistry I (every fall semester)
CHEM490/CHEM684 RNA Structure and Function (every spring semester)